Research
Gene regulatory networks are one of the main cellular infrastructures that confer defined biological function. Our research focuses on microbial synthetic biology and systems biology - the design, analysis and construction of these regulatory networks in microorganisms for cellular functionality reprogramming. These are interdisciplinary research areas at the nexus of biology, engineering and physics.
Specifically, we are interested in understanding the architecture and dynamics of naturally existing networks and exploring their relationship to cellular function. Good examples include bacterial communication networks and their engagement in cellular collective behaviors. We are also interested in building synthetic gene circuits for novel applications by assembling and editing genes and genomes inside living cells. This is very much like building integrated circuits with transistors and other elements for a computer. Along that line, microbial ecosystem engineering is of particular interest to us due to its potential for biotechnological and therapeutic purposes.
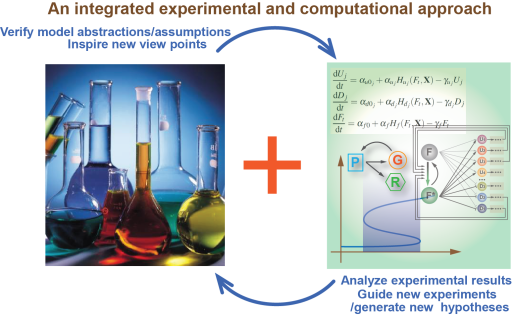
To pursue our interest, we adopt L. lactics, E. coli and other bacteria as our model organisms and employ an integrated approach that combines experiment with modeling. Experimental techniques from molecular biology and theoretical tools from nonlinear dynamics and statistical mechanics are extensively used in our research.
Our long-term goal is to uncover design principles of microbial gene regulatory networks and to apply these principles to build novel circuits for biotechnological and medical applications.